shiny中的表格
除了在shinyapp中生成图片,有时还需要生成表格,用于查看数据,以及保存下载 表格分为两类:动态和静态。静态的表格更有利于打印和输出pdf,而动态的表格提供了更多的交互式选项,还会响应屏幕的大小来进行缩放
静态表格:shiny中自带的table
shiny中自带的表格,有利于减少其他依赖包,降低内存消耗,定制和美化会受到限制
## 多文件结构的shinyapp,每个*.r代表一个R文件
## global.r## 存放一些需要运行的代码
library(tidyverse)
library(shiny)
data.main <- starwars
min_height <- unique(min(starwars$height, na.rm = TRUE))
max_height <- unique(max(starwars$height, na.rm = TRUE))
## ui.r
navbarPage("shiny::renderTable",
tabPanel("Start Narrow",
uiOutput('height_narrow_slider'), #增加滑块
tableOutput('star_narrow') #表格
),
tabPanel("Start Wide",
uiOutput('height_wide_slider'),
tableOutput('star_wide')),
tabPanel("Start List",
tableOutput('star_lists')
)
)
## server.r
function(input, output, session){
# Input widgets
output$height_narrow_slider <- renderUI({
sliderInput(
inputId = "height_limit_star_narrow",
label = "height limit",
min = min_height,
max = max_height,
value = min_height
)
})
output$height_wide_slider <- renderUI({
sliderInput(
inputId = "height_limit_star_wide",
label = "height limit",
min = min_height,
max = max_height,
value = min_height
)
})
# Table
output$star_narrow <- renderTable({
starwars %>%
select(name, species, homeworld, height) %>%
filter(height <= input$height_limit_star_narrow) %>%
arrange(desc(height))
})
# striped = TRUE,
# hover = TRUE,
# na = "[Missing]")
output$star_wide <- renderTable({
starwars %>%
select(name:species) %>%
filter(height <= input$height_limit_star_wide) %>%
arrange(desc(height))
})
output$star_lists <- renderTable({
starwars %>%
select(name, films, vehicles, starships) %>%
# unnest()
mutate_if(is.list, list(~map_chr(.,~paste(.x, collapse = "<br>"))))
}, width = "100%",
sanitize.text.function = function(x) x)
}
静态表格:kableExtra
kableExtra包的表格,也是静态的,比shiny中自带的表格具有更大的灵活性 kableExtra在server部分并不使用 render*() 函数,但ui部分还是使用tableOutput这个函数输出
说明文档链接:https://cran.r-project.org/web/packages/kableExtra/vignettes/awesome_table_in_html.html#From_other_packages
## ui.r
#install.packages("kableExtra")
library(kableExtra)
navbarPage(
"kableExtra",
tabPanel(
"Star Narrow",
fluidPage(
tableOutput('star_narrow')
)
),
tabPanel(
"Star Wide",
fluidPage(
tableOutput('star_wide')
)
),
tabPanel(
"Star List",
fluidPage(
tableOutput('star_lists')
)
),
collapsible = TRUE
)
## server.r
library(kableExtra)
library(tidyverse)
function(input, output, session){
output$star_narrow <- function(){
starwars %>%
select(name, species, homeworld, height) %>%
arrange(desc(height)) %>%
kable() %>%
kable_styling(bootstrap_options = c("striped", "hover"))
}
output$star_wide <- function(){
starwars %>%
select(name:homeworld) %>%
arrange(desc(height)) %>%
kable() %>%
kable_styling()
}
output$star_lists <- function(){
starwars %>%
select(films:starships) %>%
kable() %>%
kable_styling() ##支持list,会将list内容用逗号分隔
}
}
动态表格:Rstudio 的 DT
DT包的表格,这个属于动态表格,具有灵活交互式展现表格的特点
使用文档链接:https://rstudio.github.io/DT/
## ui.r
library(DT)
navbarPage(
"DT Interactive Tables",
tabPanel(
"Star Narrow",
fluidPage(
DTOutput('star_narrow')
)
),
tabPanel(
"Star Wide",
fluidPage(
checkboxInput("show_rownames",
label = "Show rownames?"),
DTOutput('star_wide')
)
),
tabPanel(
"Star List",
fluidPage(
DTOutput('star_lists')
)
),
collapsible = TRUE
)
## server.r
library(DT)
library(tidyverse)
function(input, output, session) {
output$star_narrow <- renderDT({
starwars %>%
select(name, species, homeworld, height) %>%
arrange(desc(height))
})
output$star_wide <- renderDT({
starwars %>%
select(name:homeworld) %>%
arrange(desc(height)) %>%
datatable(extensions = "Responsive", rownames = input$show_rownames)
})
output$star_lists <- renderDT({
starwars %>%
select(films:starships) %>%
datatable()
})
}
制作基因信息列表
## help.r
# 创建基因信息链接
# createLink for GeneCards ------------------------------------------------
geneCardsLink <- function(val,name) {
sprintf('<a href="https://www.genecards.org/cgi-bin/carddisp.pl?gene=%s" target="_blank" class="btn btn-primary">%s</a>',val,name)
}
# createLink for NCBI -----------------------------------------------------
ncbiLink <- function(val,ncbi) {
sprintf('<a href="https://www.ncbi.nlm.nih.gov/gene/?term=%s" target="_blank" class="btn btn-primary">%s</a>',val,ncbi)
}
# createLink for Esemble --------------------------------------------------
ensemblLink <- function(val,ensembl) {
sprintf('<a href="https://www.ensembl.org/Homo_sapiens/Gene/Summary?db=core;g=%s" target="_blank" class="btn btn-primary">%s</a>',val,ensembl)
}
# ui.r
fluidPage(
DT::DTOutput("gene_info")
)
# server.r
library(readr)
data <- read_csv("data/geneInfo.csv")
source("helper.R")
function(input, output, session){
geneInfo <- reactive({
tmp <- data
tmp$NCBI <- ncbiLink(tmp$SYMBOL,tmp$SYMBOL)
tmp$Ensembl.ASIA <- ensemblLink(tmp$ENSEMBL,tmp$ENSEMBL)
tmp$GeneCards <- geneCardsLink(tmp$SYMBOL,tmp$SYMBOL)
tmp
})
output$gene_info <- DT::renderDT(
#output$preview3 <- reactable::renderReactable({
DT::datatable( geneInfo(),
escape = FALSE, #决定超链接能否被渲染出来
rownames = F,
extensions = "Responsive", ##缩小页面在所在行的最前面出现加号,点击显示被隐藏的列
options=list(
pageLength = 15,
lengthMenu = list(c(15, 50, 100, -1),c(15, 50, 100, "ALL")),#-1显示所有的
scrollX = TRUE,
scrollY = TRUE,
fixedColumns = TRUE,
fixedHeader = TRUE
)
)
)
}
作业:热门的词云图+基因信息表格
# help.r
# createLink for GeneCards ------------------------------------------------
geneCardsLink <- function(val,name) {
sprintf('<a href="https://www.genecards.org/cgi-bin/carddisp.pl?gene=%s" target="_blank" class="btn btn-primary">%s</a>',val,name)
}
# createLink for NCBI -----------------------------------------------------
ncbiLink <- function(val,ncbi) {
sprintf('<a href="https://www.ncbi.nlm.nih.gov/gene/?term=%s" target="_blank" class="btn btn-primary">%s</a>',val,ncbi)
}
# createLink for Esemble --------------------------------------------------
ensemblLink <- function(val,ensembl) {
sprintf('<a href="https://www.ensembl.org/Homo_sapiens/Gene/Summary?db=core;g=%s" target="_blank" class="btn btn-primary">%s</a>',val,ensembl)
}
# ui.r
fluidPage(
sliderInput("num",
label = h3("选择热门基因的数量"),
min = 1,
max = 200,
value = c(100)),
# Copy the line below to make a set of radio buttons
radioButtons("radio", label = h3("please choose taxid"),
choices = list("human" = 9606, "pig" = 9823),
selected = 9606),
shiny::plotOutput( "cloud"),
DT::DTOutput("gene_info")
# server.r
fluidPage(
sliderInput("num",
label = h3("选择热门基因的数量"),
min = 1,
max = 200,
value = c(100)),
# Copy the line below to make a set of radio buttons
radioButtons("radio", label = h3("please choose taxid"),
choices = list("human" = 9606, "pig" = 9823),
selected = 9606),
shiny::plotOutput( "cloud"),
DT::DTOutput("gene_info")
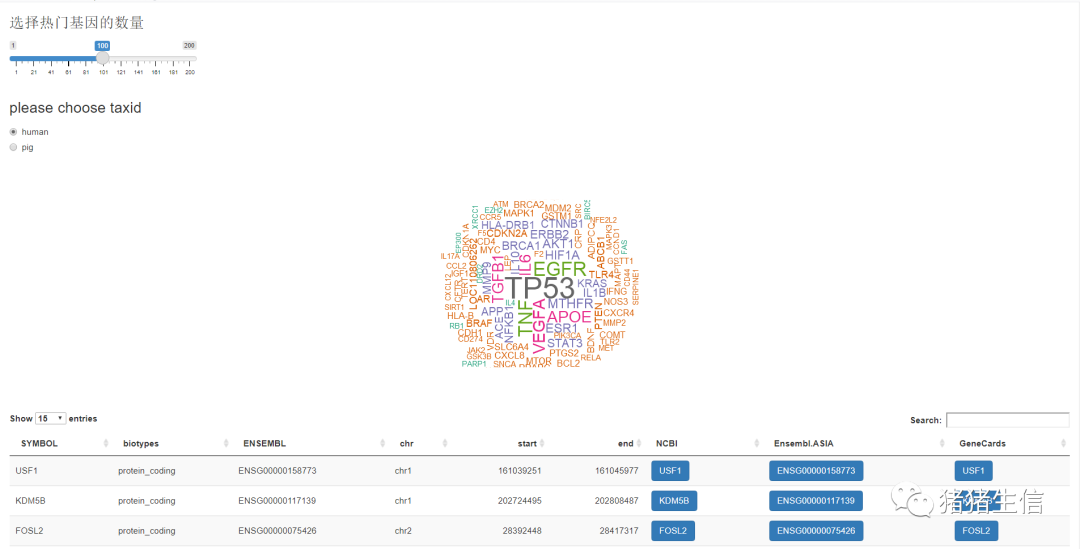
image-20210504211148970
【参考资料】
https://mp.weixin.qq.com/s/VCRukFCQaagTF6GKu2DLgA